Landmark_reference_pt_dist¶
This is a function to measure the distance from user defined points to the centroid and a point defined by the centroid coordinate along the x-axis and baseline coordinate (top of pot) along the y-axis. Calculating the vertical distance between leaf tip points to the centroid of the plant object in side-view images may provide a proxy measure of turgor pressure.
plantcv.landmark_reference_pt_dist(points_r, centroid_r, bline_r)
returns none
- Parameters:
- points_r - A list of tuples representing rescaled landmark points
- centroid_r - A tuple representing the rescaled centroid point
- bline_r - A tuple representing the rescaled baseline point
- Context:
- Used to estimate the distance and angles of landmark points relative to shape reference landmarks (centroid and pot height aka baseline)
- Data automatically gets stored into the Outputs class. Users can look at the data collected at any point during the workflow by using pcv.print_results which prints all stored data to a .json file.
- Output data stored: Summary of Output Observations
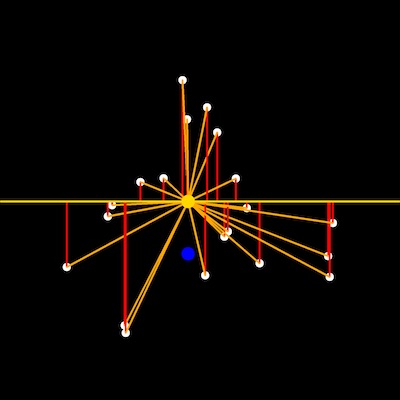
Input rescaled points, centroid and baseline points
from plantcv import plantcv as pcv
# Set global debug behavior to None (default), "print" (to file),
# or "plot" (Jupyter Notebooks or X11)
pcv.params.debug = "print"
# Identify acute vertices (tip points) of an object
# Results in set of point values that may indicate tip points
pcv.landmark_reference_pt_dist(points_r, centroid_r, bline_r)
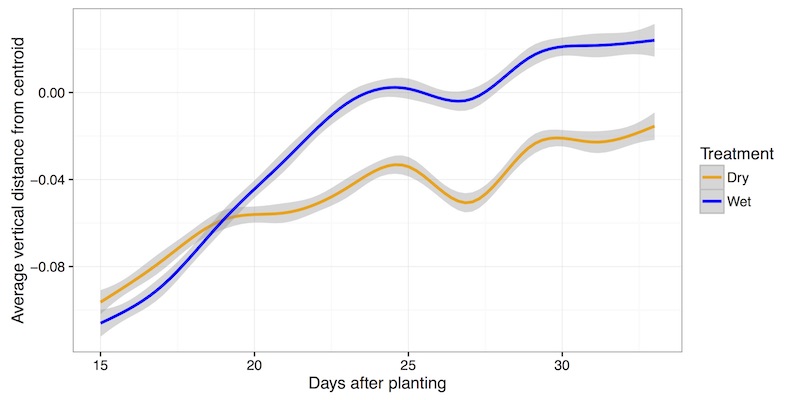
Representation of many data points collected in two treatment blocks throughout time