Analyze PSII Signal¶
Extract Fv/Fm data of objects and produce pseudocolored images.
plantcv.fluor_fvfm(fdark, fmin, fmax, mask, filename, bins=256)
returns Fv/Fm histogram headers, Fv/Fm histogram data, PSII analysis images list
- Parameters:
- fdark - image object, grayscale
- fmin - image object grayscale
- fmax - image object, grayscale
- mask - binary mask of selected contours
- filename - False or image name. If defined print image.
- bins - number of grayscale bins (0-256 for 8-bit images and 0 to 65,536), if you would like to bin data, you would alter this number (default bins=256)
- Context:
- Used to extract fv/fm per identified plant pixel.
- Generates histogram of fv/fm data.
- Generates pseudocolored output image with fv/fm values per plant pixel.
-
Example use:
-
Output Data Units:
- Bin-number - number of bins set by user
- FV/FM Bins - bin values based on number of bins set by user
- FV/FM Histogram - histogram of FV/FM ratio values for object
- FV/FM Histogram Peak - bin value of histogram peak (greatest number of pixels)
- FV/FM Median - bin value of histogram median
- F-Dark Passed QC - Check (True or False) to determine if Fdark image does not have pixel intensity values above 2000.
Fdark image

Fmin image

Fmax image

from plantcv import plantcv as pcv
# Set global debug behavior to None (default), "print" (to file), or "plot" (Jupyter Notebooks or X11)
pcv.params.debug = "print"
# Analyze Fv/Fm
fvfm_header, fvfm_data, fvfm_images = pcv.fluor_fvfm(fdark, fmin, fmax, kept_mask, filename, 1000)
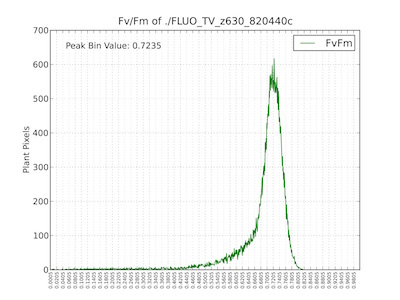
Histogram of Fv/Fm values
Pseudocolored output image based on Fv/Fm values
